
It's a polymorphism that we could use to follow the inheritance of DNA. That sequence difference doesn't necessarily mean that there's a disease associated with it. 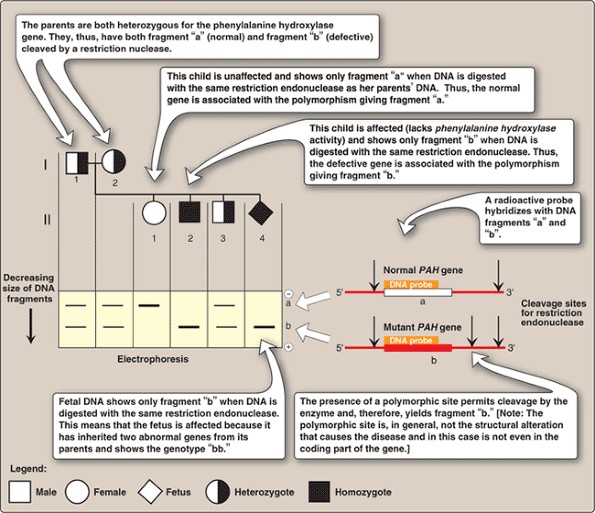
So these differences in nucleic acid sequences and restriction enzyme binding sites just mean that there's a difference in the sequence between those two people. In one person, without the enzyme site you'll see one band, and the person that has the enzyme site, you'll see two bands, representing the two cleaved products. We typically see these, or we monitor these, by isolating the DNA, cutting it with that bacterial restriction enzyme, and running it on a gel using electrophoresis. 
And that results in a polymorphism, or difference between those two people. Restriction fragment length polymorphism (RFLP) analysis utilizes restriction endonuclease digestion to identify DNA sequence polymorphisms in genes or DNA. So then, if you isolate that piece of DNA surrounding that site from two people, from one of them it will be cut by the enzyme and the other one it won't. So a single base difference between two people could result in either the presence or absence of that restriction site. Restriction Fragment Length Polymorphism (RFLP) is a technique in which organisms may be differentiated by analysis of patterns derived from cleavage of their. On this web page, you can see how RFLPs are produced and then three examples of applying RFLP analysis: paternity, disease status, and genetic mapping. So why is that useful? Well, we can take advantage of this fact to actually look for differences between people if they have that restriction enzyme site or not. RFLPs can be used to measure recombination rates which can lead to a genetic map with the distance between RFLP loci measured in centiMorgans. known as the polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) method, for the detection of single nucleotide. What is it, though? So basically, if you follow the sequence of DNA, particular sites, a series of four to eight nucleic acids, results in a restriction site where an enzyme from bacteria can actually bind and cleave that DNA. The whole protocol takes only a day to carry out. Here, we report a typical digestion protocol: add 2 L of 10× Digestion Buffer, 10 L of purified PCR products, 1 L of restriction enzyme (see Note 29), and ultrapure water up to 20 L. RFLPs have been very useful to use as markers for following a genomic DNA, either from human or other animals. In addition, a PCRrestriction fragment length polymorphism (PCRRFLP) method can be applied using restriction enzymes (2, 11, 12).
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
